top of page


Accelerating Whole-Cell mRNA Translation Simulations Using Dedicated FPGA Hardware (ACS Synthetic Biology, November 2021)
Intracellular biophysical simulations are essential for predicting how genetic modifications affect cellular resources. However, modeling the competition between thousands of mRNAs and ribosomes is computationally intensive and often creates synchronization bottlenecks that traditional parallel computing (CPUs and GPUs) cannot easily solve. We developed a novel hardware-accelerated approach using Field-Programmable Gate Arrays (FPGAs) to simulate the mRNA translation process.
1 day ago


Machine Learning-Driven Prediction and Design of Plasmid Copy Number from Replication Origin Sequences (The Royal Society Publishing, July 2025)
Precise control over Plasmid Copy Number (PCN) is a fundamental challenge in synthetic biology, directly governing the balance between recombinant protein yield and host metabolic burden. Despite its importance, computationally predicting PCN based solely on the sequence of the origin of replication (ORI) has remained an elusive goal, until now. We present a novel, interpretable machine learning framework designed to redefine plasmid engineering. By extracting over 6,000 bioc
1 day ago


Viral-Mimicry and Translational Kinetics: Advanced Modeling for Tobacco-Based Expression
In this publication published in Synthetic and Systems Biotechnology, we evaluate the sequence features that actually drive heterologous expression in tobacco. By compiling the largest known dataset of plant-based expression studies, we found that predictive accuracy increases significantly when modeling the "tRNA supply-demand" and amino acid composition rather than just host codon bias. Notably, the study reveals that successful synthetic sequences often mimic the codon usa
Apr 24


Engineering Ribosome Traffic: Improving Host Fitness and Titer via Biophysical Modeling
Standard codon optimization often ignores the "cellular cost" of protein production. In this publication, we introduce a generic computational approach to eliminate ribosomal traffic jams, a major bottleneck in protein synthesis. By engineering the kinetics of translation through synonymous mutations, we demonstrate a direct correlation between optimized ribosome allocation and improved host growth rates (fitness). Key Takeaway: Our algorithms successfully increased protein t
Apr 23


Unsupervised gene expression modeling in any given organism
Motivation: Regulation of the amount of protein that is synthesized from genes has proved to be a serious challenge in terms of analysis and prediction, and in terms of engineering and optimization, due to the large diversity in expression machinery across species. Results: To address this challenge, we developed a methodology and a software tool (ChimeraUGEM) for predicting gene expression as well as adapting the coding sequence of a target gene to any host organism. We dem
Apr 13


AI-Guided Promoter Engineering: Beyond Natural Expression Levels
AI-Driven Precision for High-Yield Engineering The Challenge: Even with advanced genetic tools, protein expression often remains suboptimal due to weak or slow-acting regulatory elements. The Innovation: The team demonstrated a breakthrough Design-Build-Test workflow. By combining biophysical modeling with synthetic biology libraries, they engineered biosensors with vastly improved sensitivity and speed. Rational Design over Trial-and-Error: Instead of random mutations, we us
Jan 11


Overcoming the challenges of genetic instability in multicopy gene design
Abstract In modern bioengineering, multicopy gene constructs are essential for reaching high protein yields and metabolic flux. However, these systems often suffer from severe genetic instability due to recombination and evolutionary pressure. Our team’s latest research, published in NAR Genomics and Bioinformatics, introduces ChimeraUGEM: a sophisticated tool that leverages biophysical modeling to design genetically stable multicopy sequences. By optimizing the "coding-regul
Dec 29, 2025

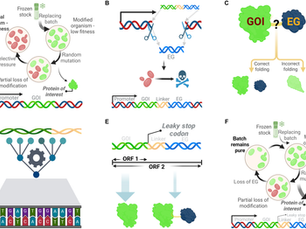
AI-directed gene fusing prolongs the evolutionary half-life of synthetic gene circuits
Abstract Evolutionary instability is a persistent challenge in synthetic biology, often leading to the loss of heterologous gene expression over time. Here, we present STABLES, a gene fusion strategy that links a gene of interest (GOI) to an essential endogenous gene (EG), with a “leaky” stop codon in between. This ensures both selective pressure against deleterious mutations and the high expression of the GOI. By leveraging a machine learning framework, we predict optimal GO
Dec 15, 2025


Deciphering plasmid replication dynamics: a computational approach to predict ColE1-like plasmid copy number
Abstract Plasmids constitute a key tool in synthetic biology, providing a versatile framework for various research and industrial ventures. An essential determinant of plasmid functionality is its copy number, impacting both protein production rates and host cell metabolic burden. However, currently there is no model that can computationally predict the plasmid copy number from the sequence of its origin of replication (ORI). We present a novel software solution tailored to s
Jul 17, 2025


Genome scale analysis of Escherichia coli with a comprehensive prokaryotic sequence-based biophysical model of translation initiation and elongation
Abstract Translation initiation in prokaryotes is affected by the mRNA folding and interaction of the ribosome binding site with the...
Jul 15, 2025


Whole cell biophysical modeling of codontRNA competition reveals novel insightsrelated to translation dynamics
Abstract The importance of mRNA translation models has been demonstrated across many fields of science and biotechnology. However, a...
Jul 15, 2025


Breakthrough in CRISPR Gene Silencing: EXPosition Tool Predicts Expression & Knockout Success
A tool for CRISPR-Cas9 sgRNA evaluation based on computational models of gene expression Background CRISPR is widely used to silence...
Dec 24, 2024


Decoding Chloroplast Translation: Novel Models and Insights for Gene Expression Engineering
Modeling the effect of rRNA-mRNA interactions and mRNA folding on mRNA translation in chloroplasts The process of translation initiation...
Dec 22, 2024


Beyond Initiation: The Role of rRNA-mRNA Interactions in Shaping Bacterial Translation Efficiency
Prokaryotic rRNA-mRNA interactions are involved in all translation steps and shape bacterial transcripts The well-established...
Dec 22, 2024


Engineering Microbiomes: Computational Frameworks for Targeted Genetic Modifications and Enhanced Biosafety
Modulating Gene Expression within a Microbiome Based on Computational Models Recent research in the field of bioinformatics and molecular...
Dec 22, 2024


Unveiling the Role of 3D Chromosomal Structure in CRISPR Efficiency and sgRNA Design
Strong association between genomic 3D structure and CRISPR cleavage efficiency CRISPR is a gene editing technology which enables precise...
Dec 22, 2024


Improved miRNA-mRNA Interaction Prediction with miBSIM: Enhanced Accuracy and Cell-Specific Insights
Advanced computational predictive models of miRNA - mRNA interaction efficiency The modeling of miRNA-mRNA interactions holds...
Dec 22, 2024


Beyond Codon Optimization: Key Determinants of Recombinant Protein Yields in Tobacco Expression Systems
Modeling coding sequence design for virus-based expression in tobacco Transient expression in Tobacco is a popular way to produce...
Dec 22, 2024


Webinar - unlocking the potential of recombinant proteins
Recap of our webinar with Dr. Guy Bushkin, director of product and synthetic biology We had an incredible session with Guy Bushkin, as...
Nov 30, 2024


Predictive Design for Stable Gene Constructs
Evolutionary Stability Optimizer (ESO) A Novel Approach to Identify and Avoid Mutational Hotspots in DNA Sequences While Maintaining High...
Jul 8, 2024
Research Powering Our Solutions
Read about our peer-reviewed publications, science and magic.
bottom of page
